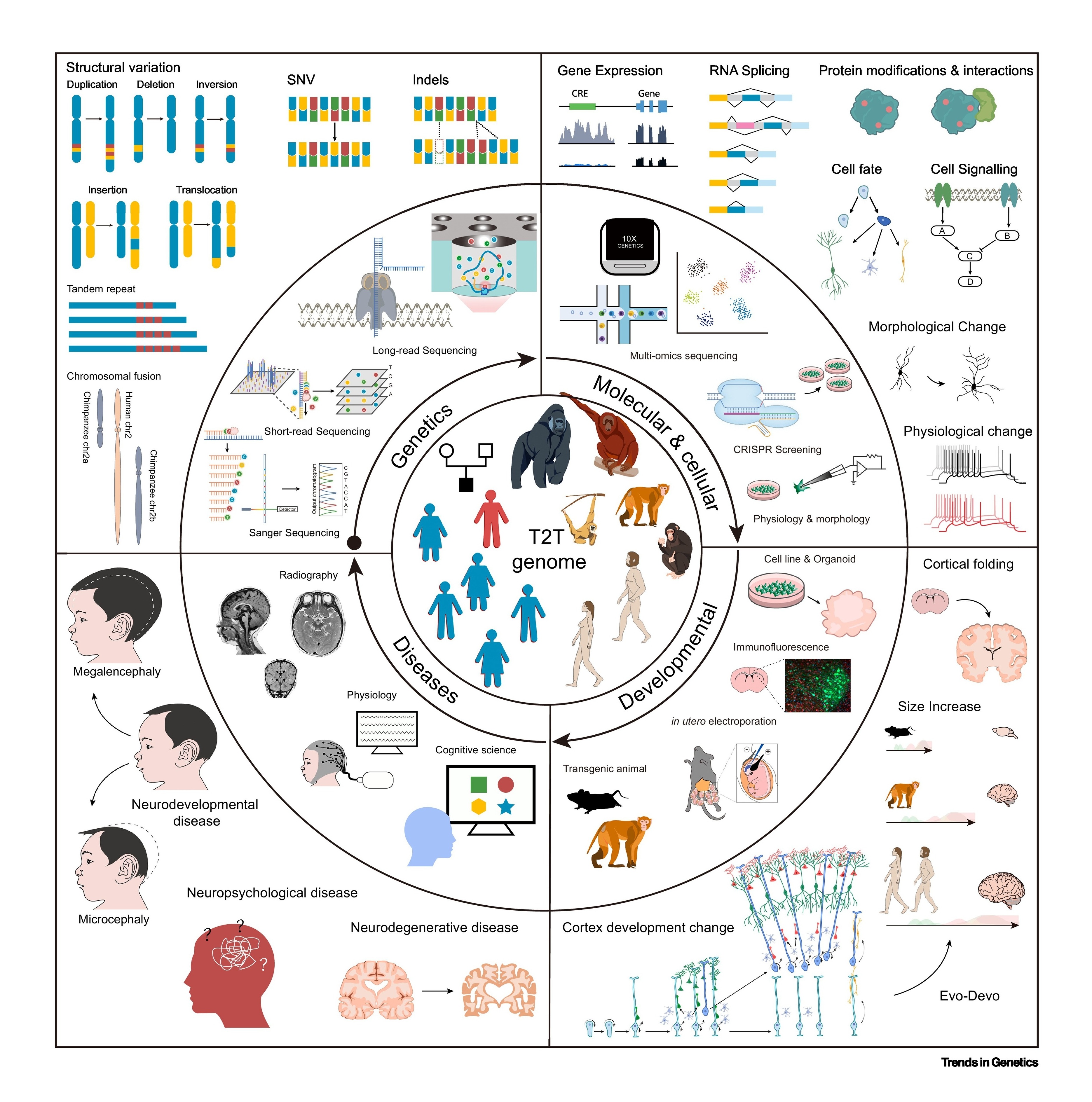
Evolution and Disease
A central goal of evolutionary biology is to understand how evolutionary processes give rise to speciation, divergence, adaptation, and biological diversity. Genetic variation—including single-nucleotide variants (SNVs), small insertions and deletions (indels), structural variants (SVs), and structurally divergent regions (SDRs)—provides the raw material for these processes and accumulates in distinct patterns across lineages. Over evolutionary time, genome architecture is further shaped by natural selection, demographic history, and neutral evolutionary forces. A fundamental open question is how genetic variation interacts with ecological context, selection, and neutrality to drive genome evolution and phenotypic innovation. Our research program addresses these questions with a particular focus on primates, combining evolutionary genomics and functional approaches to bridge evolution and disease.
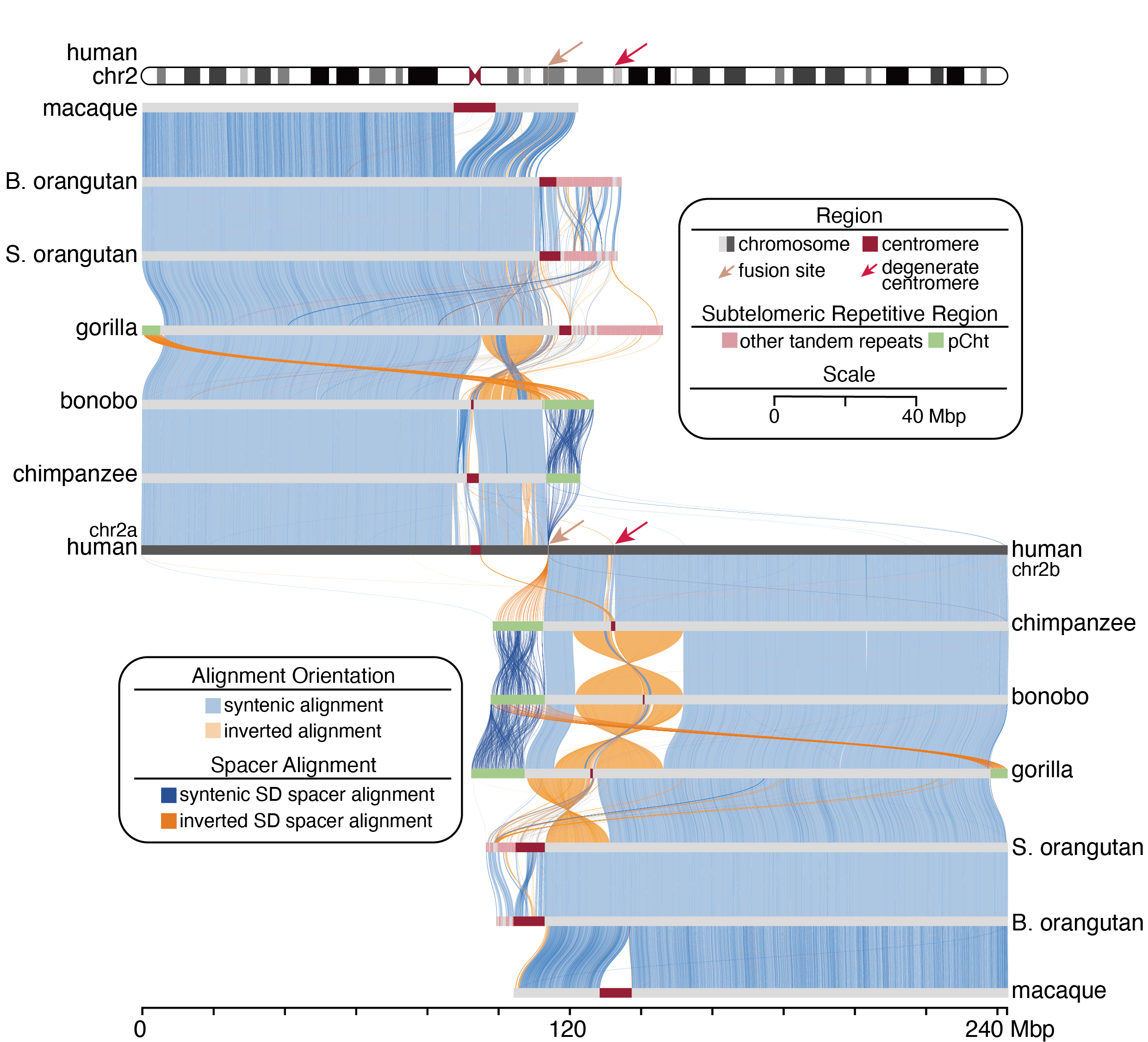

Comparative and Medical Genomics
Repetitive and structurally complex regions represent a major source of genomic variation but remain among the most challenging components of the genome to assemble and interpret. In our lab, we develop and apply computational methods to assemble complex genomic regions and to systematically characterize SVs and SDRs across primate species and populations. By integrating high-quality genome assemblies, population-scale variation data, and comparative analyses, we aim to elucidate the evolutionary origins, population dynamics, and lineage-specific patterns of SVs/SDRs. A key objective is to understand how these variants contribute to genome evolution and how they predispose or contribute to human genetic diseases.
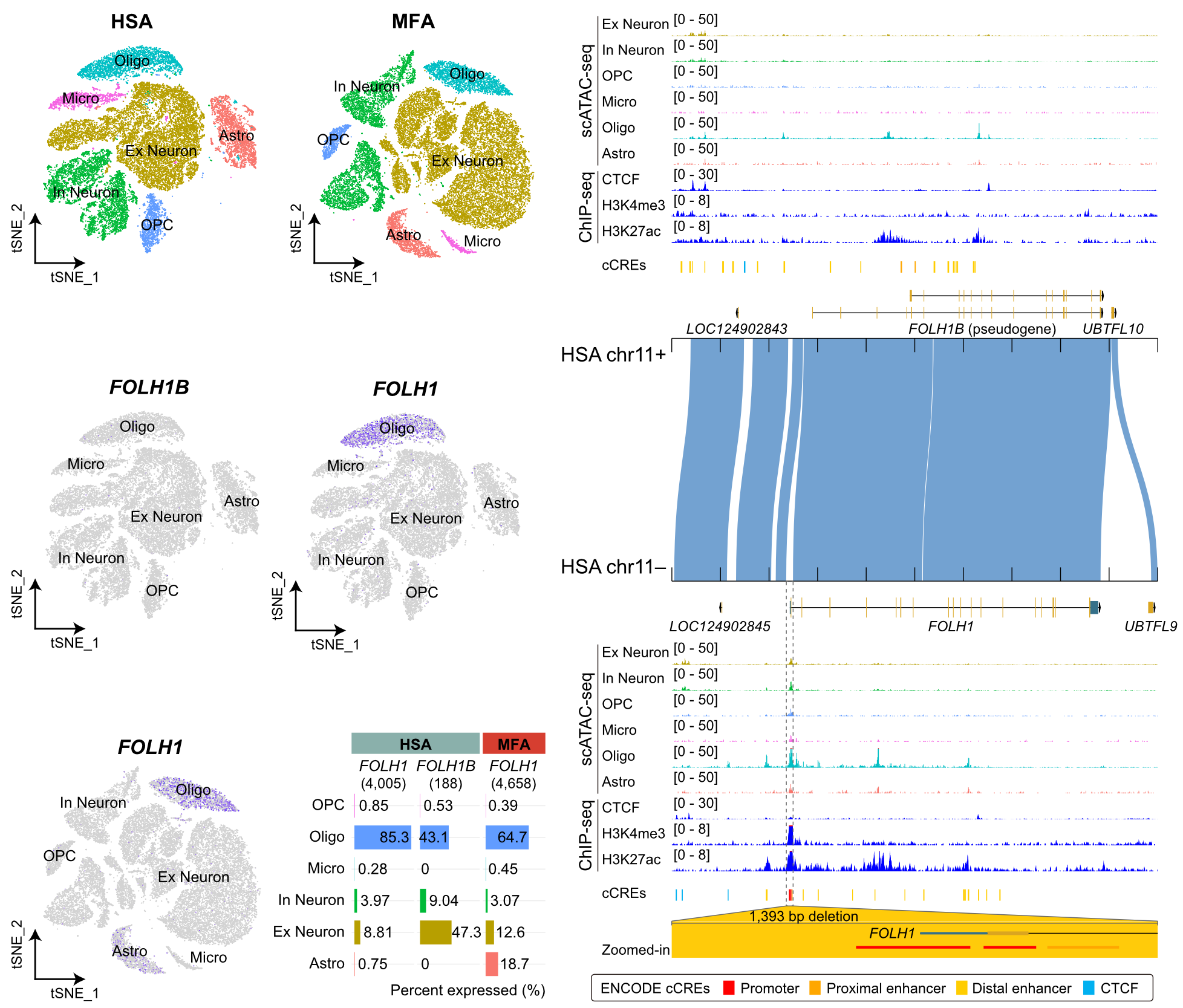
Functional Genomics of Structural Variation
Despite their evolutionary and medical relevance, the functional consequences of SVs and SDRs—particularly those underlying human-specific traits such as cortical expansion—remain poorly understood, largely due to a lack of scalable experimental approaches. Our lab addresses this gap by developing and applying high-throughput functional genomics strategies, including CRISPR-based perturbation screens, massively parallel reporter assays (MPRA), and other regulatory assays, to interrogate lineage-specific and recurrent SVs. By integrating these data with 3D genome architecture and brain organoid models, we aim to uncover the molecular and cellular mechanisms through which SV shapes gene regulation, neural development, and primate-specific traits.